It is insufficiently known that when asked, patients are very willing to share their medical information with researchers but much less so with public health authorities or third party companies. It is therefore ironic, that whereas the latter two groups do not require institutional review, researchers do. Moreover, if a study pertains to a set of patients seen at different institutions, the researcher will first have to obtain review from the institutional review boards (IRBs) of each one of the institutions. For a variety of reasons there is significant inertia working against getting one IRB to cede review of the study to another IRB. For this reason, this draft policy from the NIH is a welcome potential accelerant of biomedical research. If it is ratified, this policy will ensure that a single IRB for these multi-institutional studies becomes the rule and not the exception. A seemingly fine point to those who have never had to organize a multi-institional research study, but it might do more to advance medicine than hundreds of millions of dollars in additional research funding.
HMS - Countway Library of Medicine - Director's Blog
Harvard Medical School's Countway Library of Medicine and the new frontiers in biomedical computing
2014-12-04
2014-09-23
Worrisome Trends in Ebola
My colleague John Brownstein just shared these worrying projections for Ebola epidemiology.
2014-08-06
Near real-time tracking of Ebola
If you want to see just how Ebola is spreading (and soon regressing?) , this article provides the view and insight of how it’s done. Ten years ago, these data would percolate slowly, compiled by public health authorities and eventually appear in an academic or lay publication Now through the foundations of computational epidemiology established by Dr. Brownstein, we are able to directly monitor this in-progress disaster. Now that we have a good afferent circuit, let’s see hope the efferent circuit is even more robust.
2014-03-27
Classification system worthy of legislative intervention?
The ICD-10 diagnostic classification system is due to be adopted by US healthcare systems this year. However, new legislation would defer that change by at least a year. Who would have thought that a group of statisticians working on a list of the causes of death in 1891 would cause billions of dollars to be spent and thousands of hours invested in debating what that list should look like?
This fine system of classification that would have made Linnaeus’ head spin includes classic diagnoses such as:
V91.07xD (burn due to water-skis on fire, subsequent encounter)
W56.22xA, (struck by orca, initial encounter.)
W56.12XA (getting struck by a sea lion)
V97.33XD (Sucked into jet engine, subsequent encounter)
The orcas may have to wait another year before getting their medical recognition.
2014-03-25
Achieving clarity in cancer genomics
Approximately 3 years ago, we held a clinical bioinformatics summit to discuss with international leaders in genomics, genetic testing and clinical laboratory procedures what it would take achieve clinical grade whole genome sequencing (WGS). That is to make WGS a safe and useful component of routine diagnostic assessments in the clinic. Among the mechanisms we chose to accelerate the advent of clinical grade WGS was a competition where 3 families contributed their genomic sequence and clinical histories. Then multiple teams assembled and competed to produce the most accurate, most clinically useful diagnostic report. The competition resulted in a lot of learning and multidisciplinary team formation but also, even though we had not dared count on it, several breakthrough diagnoses. A good lay summary can be found here from Bio-IT World and an in-depth scientific summary here. Also, the cake presented by one of the mother’s to her child, who had gone for years without a diagnosis until the leading teams converged on the same mutations in the same gene, is shown below. With generous funding from Rob and Karen Hale, we kicked off the next round of the CLARITY competition as described below.
In addition to rare undiagnosed diseases, perhaps the most clinically impactful area of genomics, in the near term, is that of precision diagnostics for cancer. However, the analytic challenge is far deeper as it requires analyzing tumors that are often heterogeneous (i.e. different clones), comparing it to germ line sequence (i.e. non-tumor DNA). Then, how are we to determine which of the mutations found are relevant to the diagnosis or therapy selection? Integrating information from RNA sequence and epigenetic modifications (e.g. DNA methylation) may help and there are several good ideas being proffered by several researchers. Yet, how do we begin to bring all the data together systematically and meaningfully for patients with a malignancy? Given how early we are in this new realm of medicine, we decided to first hold a workshop where several national leaders in cancer genomics gave thoughtful assessments of the best way forward. After we we synthesize the learning from this meeting, we will announce a new CLARITY challenge for patients with recurrent cancers. Shown below are some of the speakers at the meeting which included Gad Getz, Rick Wilson, Peter Park, David Sweetser, David Margulies, Isaac Kohane, Judy Garber, Sharon Plon, Steve Chanock, Marian Harris, Heidi Rehm, Levy Garraway, and Katie Janeway. Another small but important step forward in the development of a data-driven, computationally-enabled medicine.
2014-03-03
Scholar Medical Publication DIY by Harvard Medical Students
If, as F. Scott Fitzgerald is reported to have said, action is character, then the current crop of Harvard medical students are impressive characters. Amid all the din about who controls publication, open access and shortening the loop between academia and public dissemination, a small group of determined individuals have just come out with their premiere issue of Harvard Medical Student Review. Considerable forethought, extensive deliberations with HMS leadership and creative talent have produced what is likely to become an authoritative voice for medical students and those interested in medical education and science. In the words of the student publishers themselves:
Today we are introducing the Harvard Medical Student Review, an online journal that will serve as a forum for students to participate in the conversations on healthcare and medicine. Our classmates here and around the world come from innumerable backgrounds, rich in thought-provoking experiences; we have much to share. Topics may range from research to reflections on training and experiences with patients. We hope that through writing and reflection, we can spark productive debate, generate innovation, and learn from one another at the beginning of our professional education and careers.As members of the medical community, we have a duty to review the facts and communicate the truth with clarity, precision, and humanity. In his address at the 2012 Harvard Medical School commencement ceremony, Dr. Don Berwick called upon new physicians to wield their voices proudly in carrying out this duty, as each voice “can be loud, and forceful, and confident, and will be trusted.” [2] The Harvard Medical Student Review is a space for students, trainees, and professionals to answer that call.We hope you will join us.
2014-01-24
Battle of the beauty contests
So how best to have a public debate about how we allocate the public monies for biomedical research?
.
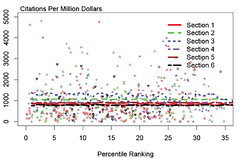
Hat tip: David Osterbur.
2014-01-23
Welcome news from the Wellcome Collection
Many of us have had the following experience. We find an interesting illustration and use it for a scholarly presentation or for a class. Then a well meaning compliance officer will notify us that if we are going to put the presentation on the web, we will have to secure the rights to the illustration for that purpose. Often the expedient response is to delete the illustration or withdraw the entire presentation from the public domain.
In this context this British invasion is a hopeful glimpse of the future. One hundred thousand pictures were just made freely available (so long as you correctly attribute their provenance). So, if you want to illustrate Harvey’s anatomical exercises of the heart and circulation, or contrast modern medical advertising to that of a well regarded phrenologist, the materials are there for you (see below), for free and without any administrative overhead. The other flank of the British open access image invasion is led by the British Library from which the charming picture below of the anatomy of the leech was taken.
2013-10-18
Help get more women into Science
There is a problem that is well articulated by one of our students, Jean Fan. To quote her:
According to the US Department of Commerce, girls (women, females, Homo sapiens with two X chromosomes, whatever term you wish to use to refer to us) remain vastly underrepresented in STEM jobs as well as among STEM degree holders. Despite filling nearly half of all jobs in the US, girls hold less than 25% of STEM jobs. And despite the rising number of girls pursuing college degrees, girls hold a disproportionally low share of STEM degrees. So what's the problem?
After articulating the problem, she provides an ingenious kickstarter project that seeks to make a small contribution to addressing this problem. I recommend you join me in supporting this project.
2013-09-28
Ghost stories for scientists
This report in Scientific American, just in time for the Halloween season, demonstrates that incorporeal beings take great interest in obesity, adipokines and leading laboratories at Harvard University. Perhaps they tired of the irreproducibility of their paranormal experiments.
Hat tip: Rachel Ramoni
2013-07-30
A thoughtful and useful report on the Aaron Swartz tragedy
This (http://swartz-report.mit.edu/docs/report-to-the-president.pdf) is a report that Professor Abelson helped author on the behalf of MIT. It is chock full of lessons for education institutions, libraries and the larger academic ecosystem, including the ancillary industries. Although section V is focused on questions for MIT, many of those same questions will find wider resonance.
2013-07-17
Innovation to grow (and track the growth of children).
I've written in the past about the bottleneck in innovation in electronic health records and how the design of such systems with substitutable "apps" would go a long way towards overcoming that bottleneck. In that context, hats off to the design team at Fjord who just won the prestigious Red Dot design award for their growth chart/growth tracking applications that are now being adopted by several vendors.
2013-06-21
Make our data work for us!
Use our data please. But for us.
2013-04-08
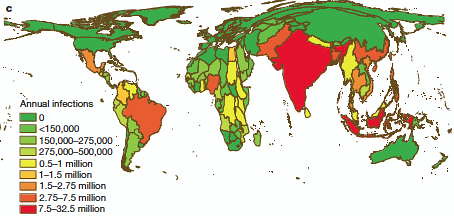
Getting Big About Mapping Dengue
Here's a very nice application of lightly used data sources about Dengue, a scourge of underdeveloped countries. As in so many areas of public health, this huge health burden is woefully under-documented. In the absence of a vetted vaccine, understanding where it is endemic is essential for the application of scarce preventive resources. This group of investigators have cleverly used a number of public but under-used data sources, including the published literature, to create a predictive map of where 390 million infections per year are occurring.
2013-01-08
Epidemic or epiphenomenon?
A number of crowd-sourced infection monitors such as FluNearYou (by our own Dr. J. Brownstein) have reported an apparent upsurge in influenza-like illnesses over the last week. The CDC has not yet reported the same trend. If the CDC then confirms this early warning, it will represent a transition from proof of principle of the citizen as health monitor (see here and here) to general public health utility. If not, then we may be witnessing an outbreak of hypochondria.
2012-12-03
Take this ontology and shove it. Or, why classification matters.
I was recently called out by one of my colleagues for saying that ontologies were boring, this despite my own doctoral work on knowledge representation. Motivating my glib comment was an image of a group of pasty-faced individuals gathered around a large boardroom table and discussing which angel fit on which pin. Events from this past weekend are a reminder why such glibness is not helpful.
The American Psychiatric Association has just approved a set of updates, revisions and changes to the reference manual (DSM5) used to diagnose mental disorders. Among the changes are those redefining the inclusion and exclusion criteria for autistic disorders. By changing which children are classified as having an autistic disorder, parents will be made to feel more or less comfortable having a child carrying the diagnosis. Just as importantly, insurance companies and school programs might shift their criteria that determine which child and family gets what kind of support and at what cost. In the near term, clinical trials for the treatment of autism may not include the same patients as they would have prior to this retaxonomization.
So, are ontologies boring? Perhaps. But they certainly belong to the class of hugely important societal constructs.
Hat tip: David Osterbur.
2012-11-20
Learning from the FDA
It is not too often that one is driven to read a report from a regulatory agency. Even rarer are the instances when we find prismatic examples of engineering and organizational leadership in these reports. That makes this strategic plan of the FDA Information Management and Office of Information Management all the more remarkable. As an inducement to read the full report, here are some results that the informatics and IT departments of many organizations, academic and industrial, would envy.
- Reducing the number of servers from 397 to 265 (by a virtualization and hosting approach).
- Not coincidentally, availability (i.e. not downtime) increased from 98.3% to 99.9996% (the difference between 30 seconds of unscheduled downtime and over 6 days unscheduled downtime).
- Billions of intrusion attempts against FDA IT Systems annually with no major information security breaches.
- Supporting annual 5-15% increases in IT capability without increases in budget.
- Training budget for personnel eliminated and training activities of personnel increased based on savings from reduced external consultant fees.
- Annual decrease in cost of data storage.
2012-10-23
No Publication Without Taxation?
I recently obtained a copy of a presentation by Elsevier representatives describing how they price their publications for different academic (i.e. university) customers. They describe how they place each institution into one of a small number of pricing tiers. How do they decide this categorization? A major component of the decision is the research intensity of the institution to which they were selling access to their publications. What was the measure of research intensity? Publication volume as measured by the SCOPUS database (an Elsevier product).
To better understand this policy, I contacted Elsevier and spoke by phone with a very cordial and clear executive. I asked why it seemed that the more we published, the more we were going to be charged by Elsevier to read these publications?
He explained that research intensity served as a proxy for the value that institutions might place on these publications. After all, if we value these publications, then we are more likely to download/use them. And pricing should reflect value.
I then asked why they were not simply measuring the download rate for each journal to directly charge for usage rather than using a proxy measure. The executive explained that their advisory board had recommended not to use the download rate as that might reduce usage thereby impeding scholarship.
So solicitous of the publisher. I knew it would be futile to bring up the effect on scholarship of inflation rates of publication prices that would make the rate of medical care inflation appear flat by comparison. A continual record of inflation that reduces every year the fraction of their own scholarly output made available to scholars throughout the world's academic institutions.
2012-09-03
Yet Another Healthcare Research Steeplechase Barrier
I was recently informed that I have to take an on-line course about conflict of interest and then document all commercial activities. I was then informed of the same duty three more times. Because I am engaged in research at multiple universities, hospitals and medical schools, these education and reporting activities have to be done multiple times and of course the forms and "educational" syllabi are all different.
Like all large, heavily funded (and rewarded) organized activities, healthcare research has acquired over time some practices that originally made sense but have evolved into anti-productive structures. There have been a number of highly publicized examples of truly egregious behavior by researchers who have hidden their conflicts of interest. Individuals who used the podium of academia and the cachet of their presumed impartiality to declaim, opine, publish regarding a device, therapy, or diagnostic procedure where they stood to gain financially. For decades, it was made abundantly clear to all investigators that any such conflict was to be disclosed in grant applications, in publications and presentations. Yet. it was left up the the investigator to decide whether or not there was indeed a conflict. This ultimately became an all too carefully parsed taxonomical challenge and rather than waste time in such parsing, several of us went the route of full transparency. For example, I have listed on the web all my commercial activities so that my colleagues and the public can decide whether or not there is indeed a conflict.
Presumably because not everyone has traversed this route of transparency, there are now a host of new regulations for annual disclosure. Some of them appear obvious (e.g. disclosure of equity ownership or payment for speaking), others less so (e.g reimbursement for cab fare to attend a commercial conference where there are no speaking fees). In the end these disclosures are important and yet are each of the healthcare research institutions so different as to require different reporting and educational mechanisms? It seems there is an opportunity for an enterprising company or apparatchik to create a single, authoritative form and set of educational materials. Let's just hope they keep it simple.
2012-08-16
Hungry for DNA Games?
Thirty teams world-wide are apparently hungry enough and willing to contribute to making genomic medicine possible. Their efforts will help reduce to practice a game that to date is only within reach of star teams. May the odds be ever in our favor.